
| Size | Price | Stock | Qty |
|---|---|---|---|
| 1mg |
|
||
| 2mg |
|
||
| 5mg |
|
||
| 10mg |
|
||
| 25mg |
|
||
| 50mg |
|
||
| 100mg |
|
||
| 250mg |
|
||
| 500mg | |||
| Other Sizes |
Purity: ≥98%
MM-102 (also known as HMTase Inhibitor IX) is a novel and potent peptidomimetic inhibitor of MLL1 (Mixed lineage leukemia 1) with anticancer activity. It inhibits MLL1 with an IC50 of 0.4 μM in a cell-free assay. MM-102 inhibited the growth of leukemia cells harboring MLL1 fusion proteins.
| Targets |
MLL1-WDR5 PPI (IC50 = 2.4 nM)
MLL1-WDR5 protein-protein interaction (Ki = ~1.2 nM determined by isothermal titration calorimetry (ITC) for the interaction between recombinant MLL1 SET domain (residues 3749–3969) and WDR5; IC₅₀ = ~5 nM in AlphaScreen binding assay; no significant binding to other WDR5-interacting proteins (e.g., Ash2L) with Ki > 10 μM, confirming selectivity) [1] - MLL1-WDR5 protein-protein interaction (IC₅₀ = ~6 nM in HTRF binding assay for MLL1-WDR5 complex; inhibits MLL1-mediated histone H3K4 methylation in renal tubular cells, with no off-target effects on EZH2 or DOT1L) [2] |
|---|---|
| ln Vitro |
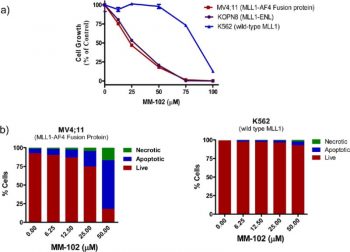
In vitro activity: MM-102, as a MLL1 mimetic, shows high binding affinities to WDR5 with IC50 of 2.9 nM and Ki of < 1 nM. In the MLL1-AF9 transduced murine cells, MM-102 specifically reduces expression of two critical MLL1 target genes (HoxA9 and Meis-1), which are required for MLL1 mediated leukemogenesis. In addition, MM-102 effectively and selectively inhibits cell growth and induces apoptosis in leukemia cells harboring MLL1 fusion proteins
Kinase Assay: The HMT assay is performed in 50 mM HEPES pH 7.8, 100 mM NaCl, 1.0 mM EDTA, and 5% glycerol at 22 °C. Each reaction contains 1.5 μCi of the co-factor, 3H-S-adenosylmethionine. H3 10-residue peptide is used as the substrate at 50 μM. Compounds are added at concentrations ranging from 0.125 to 128 μM and incubated with the pre-assembled WDR5/RbBP5/ASH2L complex at a final concentration of 0.5 μM for each protein for 2–5 min. Reactions are initiated by addition of the MLL1 protein at a final concentration of 0.5 μM and allowed to proceed for 30 min before preparing scintillation counting. To count samples, reactions are spotted on separate squares of P81 filter paper and precipitated by submerging in freshly prepared 50 mM sodium bicarbonate buffer with pH 9.0. After washing and drying, samples are vortexed in Ultima Gold scintillation fluid and counted. As a negative control, assays are performed using 0.5 μM MLL1/WDR5/RbBP5/ASH2L complex assembled with the non-interacting mutant, WDR5D107A. Cell Assay: MM-102 significantly inhibited the expression of HoxA9 and Meis-1, two critical MLL1 target genes in MLL1 fusion protein mediated leukemogenesis. MM-102 (6.25, 25, 50 μM) also specifically suppressed cell growth and induced apoptosis in leukemia cells harboring MLL1 fusion proteins. MV4;11, KOPN8, and K562 cells are cultured in RPMI 1640 medium (ATCC) supplemented with 10% fetal bovine serum and 100 U/L penicillin-streptomycin and incubated at 37 °C under 5% CO2. Cells are seeded into 12-well plates for suspension at a density of 5 × 105 per well (1 mL) and treated with either vehicle control (DMSO, 0.2%) or MM-102 for 7 days. The medium is changed every 2 days, and compounds are resupplied. The CellTiter-Glo Luminescent Cell Viability Assay kit is used following the manufacturer’s instruction. First, 100 μL of the assay reagent is added into each well, and the content is mixed for 2 min on an orbital shaker to induce cell lysis. After 10 min incubation at room temperature, the luminescence is read on a microplate reader 1. Binding affinity and selectivity (leukemia focus): MM-102 specifically binds to the MLL1-WDR5 interaction interface. ITC experiments showed a 1:1 binding stoichiometry between MM-102 and the MLL1-WDR5 complex, with a Ki of ~1.2 nM. AlphaScreen assays confirmed it inhibited MLL1-WDR5 binding with an IC₅₀ of ~5 nM, while showing no binding to WDR5-Ash2L (Ki > 10 μM) or WDR5-RbBP5 (Ki > 15 μM) [1] 2. Inhibition of MLL1 methyltransferase activity: MM-102 (0.1–10 nM) dose-dependently inhibited MLL1-mediated H3K4 trimethylation (H3K4me3) in a reconstituted PRC1-like complex (MLL1-WDR5-Ash2L-RbBP5). At 5 nM, H3K4me3 was reduced by ~80% (Western blot). It had no effect on other histone methyltransferases (e.g., EZH2, G9a) even at 1 μM [1] 3. Antiproliferative activity in MLL-rearranged leukemia cells: MM-102 suppressed proliferation of MLL-rearranged leukemia cell lines: MV4-11 (MLL-AF4) had an IC₅₀ of ~0.8 μM, RS4;11 (MLL-AF4) had an IC₅₀ of ~1.0 μM. Non-MLL-rearranged leukemia cells (K562) were less sensitive (IC₅₀ > 10 μM). Treatment with 1 μM MM-102 for 72 h reduced MV4-11 cell viability by ~75% (MTT assay) [1] 4. Downregulation of MLL target genes in leukemia: MM-102 (0.5–2 μM for 48 h) dose-dependently reduced H3K4me3 levels at the HOXA9 and MEIS1 promoters (ChIP-qPCR: ~65% reduction at 1 μM) in MV4-11 cells. qRT-PCR showed HOXA9 mRNA (-60%) and MEIS1 mRNA (-55%) downregulation, while Western blot confirmed ~50% reduction in HOXA9 protein [1] 5. Inhibition of cisplatin-induced renal tubular cell apoptosis: MM-102 (1–5 μM) protected HK-2 human renal tubular cells from cisplatin (20 μM)-induced apoptosis. Annexin V-FITC/PI staining showed apoptotic rates decreased from ~45% (cisplatin alone) to ~12% (cisplatin + 3 μM MM-102). Western blot detected reduced cleaved caspase-3 (~70% reduction) and cleaved PARP (~65% reduction) [2] 6. Regulation of E-cadherin and p53 in renal cells: MM-102 (3 μM for 48 h) reversed cisplatin-induced downregulation of E-cadherin in HK-2 cells (2.5-fold increase in protein levels, Western blot). It also reduced cisplatin-induced p53 activation: phospho-p53 (Ser15) levels were reduced by ~60%, and p53 target pro-apoptotic gene Bax mRNA was downregulated by ~50% (qRT-PCR) [2] |
| ln Vivo |
MM-102 attenuates AKI after cisplatin administration in mice[2]
To investigate the role of MLL1/WDR5 in cisplatin-induced AKI, mice were treated with MM102, an inhibitor of the MLL1/WDR5 complex, or vehicle 2 h before cisplatin administration (20 mg/kg, intraperitoneally injection). MM102 was then given daily for three consecutive days. Blood samples and kidney tissue were collected 72 h after cisplatin injection. Blood urea nitrogen (BUN) and serum creatinine (SCr) were used as measures of renal function. As shown in Fig. 1A, BUN levels in cisplatin group were much higher than that in control group (6.217 ± 0.374 vs. 2.420 ± 0.470 mmol/L) (***P < 0.001); MM102 treatment reduced the cisplatin-boosted BUN to 3.172 ± 0.114 mmol/L (**P < 0.01). Similarly, SCr was 68.126 ± 10.217 μmol/L in cisplatin-alone group (Fig. 1B), higher than that in the control group (10.322 ± 2.135 μmol/L) (**P < 0.01); MM102 treatment significantly reduced SCr to 20.922 ± 4.016 μmol/L (**P < 0.01); MM102 alone had little effect on either BUN or SCr. MM-102 reduces apoptosis, along with reduced p53 phosphorylation and retained E-cadherin expression in vivo[2] IF staining indicated that neutrophil gelatinase-associated lipocalin (NGAL, an early biomarker of AKI) was increased in kidneys exposed to cisplatin relative to sham-operated kidneys. Administration of MM102 dramatically reduced NGAL expression in cisplatin-injured kidneys (Fig. 2A, B). Consistently, TdT-mediated dUTP-X nick-end labeling (TUNEL) staining displayed increased number of apoptotic cells in injured kidney and MM102 largely inhibited this response (Fig. 2A, C). Moreover, increased expression of NGAL and cleavage of caspase-3 (C-cas3, a recognized marker of apoptosis) in the kidney after cisplatin administration were detected by immunoblot analysis; treatment with MM102 returned these changes to base levels. 1. Tumor growth inhibition in MLL-rearranged leukemia xenografts: In NOD/SCID mice bearing MV4-11 (MLL-AF4) subcutaneous xenografts, MM-102 was administered by oral gavage at 20 mg/kg once daily for 21 days. Day 21 tumor volume: ~250 mm³ (treatment) vs. ~820 mm³ (vehicle), tumor growth inhibition rate (TGI) = ~69%. Tumor weight at sacrifice: ~100 mg (treatment) vs. ~340 mg (vehicle), ~71% reduction. Tumor tissues showed ~60% lower H3K4me3 and ~55% lower HOXA9 protein (Western blot) [1] 2. Survival prolongation in disseminated leukemia: In a lethal MV4-11 intravenous xenograft model, MM-102 (20 mg/kg, oral gavage, qd for 21 days) prolonged median survival from 22 days (vehicle) to 33 days. At day 38, 25% of treated mice survived vs. 0% vehicle [1] 3. Protection against cisplatin-induced acute kidney injury (AKI): C57BL/6 mice were pre-treated with MM-102 (10 mg/kg, intraperitoneal injection) 1 h before cisplatin (20 mg/kg, intraperitoneal injection). On day 7 post-cisplatin, MM-102 reduced serum creatinine (from 180 μmol/L to 95 μmol/L) and blood urea nitrogen (BUN, from 35 mmol/L to 18 mmol/L) (key AKI markers). Renal histopathology showed reduced tubular necrosis (tubular injury score: 1.2 vs. 3.8 in cisplatin alone) and fewer TUNEL-positive apoptotic cells (~70% reduction) [2] 4. Renal molecular changes in AKI model: Kidney tissues from MM-102-treated mice had increased E-cadherin protein (+2.3-fold) and decreased phospho-p53 (-65%) and cleaved caspase-3 (-70%) (Western blot). MLL1 target gene Hoxa9 mRNA was downregulated by ~50% (qRT-PCR), confirming on-target inhibition [2] |
| Enzyme Assay |
Competitive Binding Assay[1]
Binding affinities of all the synthesized compounds were determined using a fluorescence-polarization (FP)-based competitive binding assay; the details of this assay have been described earlier. In Vitro Histone Methyltransferase (HMT) Assay[1] The HMT assay was performed in 50 mM HEPES pH 7.8, 100 mM NaCl, 1.0 mM EDTA, and 5% glycerol at 22 °C. Each reaction contained 1.5 μCi of the co-factor,3H-S-adenosylmethionine. H3 10-residue peptide was used as the substrate at 50 μM. Compounds were added at concentrations ranging from 0.125 to 128 μM and incubated with the pre-assembled WDR5/RbBP5/ASH2L complex at a final concentration of 0.5 μM for each protein for 2–5 min. Reactions were initiated by addition of the MLL1 protein at a final concentration of 0.5 μM and allowed to proceed for 30 min before preparing scintillation counting. To count samples, reactions were spotted on separate squares of P81 filter paper (Whatman) and precipitated by submerging in freshly prepared 50 mM sodium bicarbonate buffer with pH 9.0. After washing and drying, samples were vortexed in Ultima Gold scintillation fluid and counted. As a negative control, assays were performed using 0.5 μM MLL1/WDR5/RbBP5/ASH2L complex assembled with the non-interacting mutant, WDR5D107A. 1. ITC for MLL1-WDR5 binding: Recombinant human WDR5 (20 μM) and MLL1 SET domain (10 μM) were mixed in buffer (20 mM Tris-HCl pH 7.5, 150 mM NaCl, 1 mM DTT) to form the MLL1-WDR5 complex. MM-102 (50 μM in the same buffer) was titrated into the complex at 25°C. Heat changes were recorded, and data were fitted to a 1:1 binding model to calculate the Ki [1] 2. AlphaScreen MLL1-WDR5 inhibition assay: Biotinylated MLL1 peptide (residues 3765–3785, 50 nM) and GST-tagged WDR5 (50 nM) were incubated with serial concentrations of MM-102 (0.1 nM–100 nM) in assay buffer (50 mM HEPES pH 7.4, 100 mM NaCl, 0.1% BSA) for 1 h at room temperature. Anti-biotin donor beads and anti-GST acceptor beads were added, and fluorescence was measured at 615 nm. IC₅₀ was derived from dose-response curves [1] 3. MLL1 methyltransferase activity assay: Reconstituted MLL1 complex (MLL1-WDR5-Ash2L-RbBP5, 10 nM each) was incubated with histone H3 (1–21) peptide (2 μM), S-adenosyl-L-methionine (SAM, 10 μM), and MM-102 (0.1 nM–10 μM) at 37°C for 2 h. H3K4me3 levels were detected by ELISA using anti-H3K4me3 antibody. Percentage inhibition was calculated relative to vehicle, and IC₅₀ was determined [1] 4. HTRF MLL1-WDR5 binding assay (renal focus): Fluorescein-labeled MLL1 peptide (50 nM) and WDR5 (50 nM) were incubated with MM-102 (0.1 nM–100 nM) in HTRF buffer (50 mM Tris-HCl pH 7.5, 100 mM NaCl) for 1 h. Anti-fluorescein cryptate and anti-WDR5 d2 antibodies were added, and time-resolved fluorescence was measured. IC₅₀ was calculated from the dose-response curve [2] |
| Cell Assay |
qRT-PCR Analysis of HOXA9 and MEIS-1 Genes[1]
Murine MLL1-AF9 transformed bone marrow cells were obtained by transducing normal murine bone marrow cells with MLL1-AF9 oncogene according to the procedures described by Tan et al.22 MM-102 and C-MM-102 were dissolved in DMSO. The transformed cells were treated with MM-102 (25 μM, 50 μM), C-MM-102 (50 μM), and Mock (0.2% DMSO), giving a final concentration of 0.2% DMSO in all the samples. Total RNA was isolated from MLL1-AF9 transduced mouse bone marrow cells after 96 h treatment using Trizol and the RNEASY kit according to the protocol described earlier.23 The cDNA was generated using random priming with the SuperScript III kit. Real-time PCR amplifications of HoxA9, Meis1, and GAPDH genes were carried out with primers specific for each gene in the presence of SYBR dye. Relative quantification of each gene transcript was carried out as described in our previous work.10 The results were presented as relative expression to Mock treatment after normalizing to an internal loading control (e.g., GAPDH or total input RNA). Cell Growth and Apoptosis Studies of Leukemia Cell Lines[1] MV4;11, KOPN8, and K562 cells were a generous gift from Dr. Jolanta Grembecka (University of Michigan). MV4;11, KOPN8, and K562 cells were cultured in RPMI 1640 medium (ATCC) supplemented with 10% fetal bovine serum and 100 U/L penicillin-streptomycin and incubated at 37 °C under 5% CO2. Cells were seeded into 12-well plates for suspension at a density of 5 × 105 per well (1 mL) and treated with either vehicle control (DMSO, 0.2%) or MM-102 for 7 days. The medium was changed every 2 days, and compounds were resupplied. 1. MTT antiproliferation assay (leukemia): MV4-11 or RS4;11 cells were seeded in 96-well plates (3×10³ cells/well) and cultured overnight. Serial concentrations of MM-102 (0.01 μM–20 μM) were added, and cells were incubated for 72 h (37°C, 5% CO₂). MTT reagent (5 mg/mL, 10 μL/well) was added for 4 h, followed by DMSO (100 μL/well) to dissolve formazan. Absorbance at 570 nm was measured, and IC₅₀ was calculated via nonlinear regression [1] 2. Western blot for epigenetic and apoptotic markers (leukemia): MV4-11 cells were treated with MM-102 (0.5–2 μM) for 48 h. Nuclear proteins were extracted for H3K4me3/HOXA9 detection, and total proteins for apoptotic markers. Samples were separated by SDS-PAGE, transferred to PVDF membranes, and probed with primary antibodies (anti-H3K4me3, anti-HOXA9) and HRP-conjugated secondary antibodies. Signals were quantified relative to GAPDH [1] 3. ChIP-qPCR for H3K4me3 at target promoters: MV4-11 cells treated with 1 μM MM-102 for 48 h were cross-linked with 1% formaldehyde. Chromatin was sheared by sonication, incubated with anti-H3K4me3 antibody, and pulled down with protein A/G beads. DNA was purified and analyzed by qPCR using primers for HOXA9/MEIS1 promoters [1] 4. Apoptosis assay (renal tubular cells): HK-2 cells were pre-treated with MM-102 (1–5 μM) for 2 h, then exposed to cisplatin (20 μM) for 48 h. Cells were stained with Annexin V-FITC and PI for 15 min (room temperature, dark) and analyzed by flow cytometry. Apoptotic cells (Annexin V⁺/PI⁻ + Annexin V⁺/PI⁺) were counted [2] 5. Western blot for renal cell markers: HK-2 cells were treated as above, and total proteins were extracted. Membranes were probed with anti-E-cadherin, anti-phospho-p53 (Ser15), anti-cleaved caspase-3, and anti-GAPDH antibodies. Band intensities were quantified using ImageJ software [2] 6. qRT-PCR for renal gene expression: Total RNA from treated HK-2 cells was extracted with TRIzol, reverse-transcribed to cDNA, and qRT-PCR was performed with primers for Bax, Hoxa9, and GAPDH. Relative mRNA levels were calculated via 2^(-ΔΔCt) [2] |
| Animal Protocol |
Animals models of AKI and treatment[2]
Male C57BL/6J mice aged 6–8 weeks and weighing 20–25 g were purchased from the Jackson Laboratory. The mice were randomly divided into four groups: (1) control, (2) MM-102, (3) cisplatin, and (4) MM-102 plus cisplatin. Cisplatin was intraperitoneally injected at the dose of 20 mg/kg. MM-102 (15 mg/kg) dissolved in solvent containing 10% DMSO and 90% corn oil was administered intraperitoneally 2 h before the cisplatin injection and then given daily for three consecutive days. The dose of MM-102 was selected according to a previous report. For the control and cisplatin-alone groups, mice were injected with an equivalent amount of solvent. Mice in the control and MM-102 groups were injected with an equal volume of a normal saline solution. All the mice were euthanized 72 h after cisplatin injection. Blood samples and kidney tissues were collected for further analysis. All experimental protocols were performed according to the National Institutes of Health Guidelines on the Care and Use of Laboratory Animals and approved by the Lifespan Animal Welfare Committee. The authorization number for the use of laboratory animals is 5074-19. 1. Leukemia subcutaneous xenograft model: Female NOD/SCID mice (6–8 weeks old) were subcutaneously injected with 5×10⁶ MV4-11 cells (PBS:Matrigel = 1:1) into the right flank. When tumors reached 100–150 mm³, mice were randomized into vehicle (n=6) and MM-102 (n=6) groups. MM-102 was dissolved in DMSO:PEG400:0.9% saline (15:35:50, v/v/v) to 4 mg/mL. Mice received 20 mg/kg MM-102 via oral gavage once daily for 21 days; vehicle received the same volume of solvent. Tumor volume (length × width² / 2) and body weight were measured every 3 days. Tumors were collected at sacrifice for molecular analysis [1] 2. Leukemia disseminated xenograft model: Female NOD/SCID mice were intravenously injected with 2×10⁶ MV4-11 cells via tail vein. Three days later, mice were randomized into vehicle (n=8) and MM-102 (n=8) groups. MM-102 was administered as above (20 mg/kg, oral gavage, qd) for 21 days. Mice were monitored for morbidity (weight loss >20%, lethargy), and survival time was recorded [1] 3. Cisplatin-induced AKI mouse model: Male C57BL/6 mice (8–10 weeks old) were divided into 3 groups (n=6/group): control, cisplatin alone, cisplatin + MM-102. MM-102 was dissolved in DMSO:corn oil (10:90, v/v) to 2 mg/mL. Mice in the combination group received 10 mg/kg MM-102 via intraperitoneal injection 1 h before cisplatin (20 mg/kg, intraperitoneal injection). Control mice received vehicle. On day 7 post-cisplatin, blood was collected for creatinine/BUN measurement, and kidneys were harvested for histopathology and Western blot [2] |
| ADME/Pharmacokinetics |
1. Oral bioavailability in mice: Female CD1 mice were administered MM-102 orally (20 mg/kg) orally or intravenously (5 mg/kg). Blood samples were collected at 0.25, 0.5, 1, 2, 4, 8, and 24 hours post-administration. The concentration of MM-102 in plasma was determined by liquid chromatography-tandem mass spectrometry (LC-MS/MS). The oral bioavailability was calculated to be ~42% (oral AUC₀₋∞ / intravenous AUC₀₋∞ × intravenous dose / oral dose × 100%) [1]
2. Plasma pharmacokinetics (oral): After CD-1 mice were orally administered 20 mg/kg MM-102, the key parameters were: Cₘₐₓ = ~3.1 μM, Tₘₐₓ = ~1.2 h, t₁/₂ = ~3.5 h, AUC₀₋₂₄ₕ = ~10.8 μM·h [1] 3. Tissue distribution: NOD/SCID tumor-bearing mice (MV4-11 xenograft tumor) were administered 20 mg/kg MM-102 by gavage. 1.2 hours after administration (Tₘₐₓ), tissue samples were collected and analyzed by LC-MS/MS. The concentrations were: tumor tissue ≈ 2.8 μM, liver ≈ 4.5 μM, spleen ≈ 3.8 μM, lung tissue ≈ 2.2 μM, and kidney ≈ 1.9 μM. The tumor tissue concentration exceeded the in vitro IC₅₀ value of MV4-11 cells (0.8 μM) [1]. 4. Kidney tissue distribution (AKI model): C57BL/6 mice were intraperitoneally injected with 10 mg/kg MM-102. 1 hour after administration, the concentration of MM-102 in the kidney tissue was approximately 1.5 μM (LC-MS/MS), which was sufficient to inhibit MLL1-WDR5 interaction [2]. |
| Toxicity/Toxicokinetics |
1. Acute toxicity in mice: Female CD1 mice were administered MM-102 orally at doses of 50, 100, 150 and 200 mg/kg. No death or significant toxic reactions (e.g., weight loss, lethargy) were observed at the 200 mg/kg dose. The LD₅₀ was determined to be >200 mg/kg [1]
2. Chronic toxicity in leukemia model: In a 21-day gavage study (20 mg/kg), mice treated with MM-102 did not show significant weight loss (maximum change: 5% decrease compared to the carrier group). Serum biochemical indicators (ALT, AST, creatinine, urea) were normal, and hematological indicators (white blood cells, red blood cells, platelets) were normal [1] 3. AKI model toxicity: Mice treated with MM-102 (10 mg/kg, intraperitoneal injection) in combination with cisplatin did not show additional hepatotoxicity compared to cisplatin alone. Serum ALT/AST levels were normal, and no MM-102-induced damage was observed in kidney tissue pathology. No changes in body weight or behavior were observed [2]. 4. Plasma protein binding: MM-102 (1 μM) was incubated with mouse plasma at 37°C for 1 hour. Unbound drug was separated by ultrafiltration (30 kDa molecular weight cutoff) and measured by LC-MS/MS. The plasma protein binding rate was approximately 93% [1][2]. |
| References |
|
| Additional Infomation |
Mixed-lineage leukemia 1 (MLL1) is a histone H3 lysine 4 (H3K4) methyltransferase, and targeting MLL1 enzyme activity has been proposed as a novel therapeutic strategy for acute leukemia carrying MLL1 fusion proteins. The MLL1/WDR5 protein-protein interaction is crucial for MLL1 enzyme activity. In this study, we designed a large number of peptide mimics targeting the MLL1/WDR5 interaction based on the minimum binding motif of MLL1, -CO-ARA-NH-. Our study designed high-affinity peptide mimics that exhibited potent antagonism of MLL1 activity in a fully recombinant in vitro H3K4 methyltransferase activity assay with a binding affinity (Ki < 1 nM) to WDR5. Cocrystal structure determination of two potent peptide mimics with WDR5 revealed the structural basis for their high-affinity binding to WDR5. Evaluation of one of the peptide mimics, MM-102, in bone marrow cells transfected with the MLL1-AF9 fusion construct showed that the compound effectively reduced the expression of HoxA9 and Meis-1, two key target genes in the MLL1 fusion protein-mediated leukemia process. MM-102 also specifically inhibited the growth of leukemia cells carrying the MLL1 fusion protein and induced their apoptosis. Our study is the first to validate the concept of designing small molecule inhibitors of WDR5/MLL1 protein-protein interaction as a novel treatment for acute leukemia carrying the MLL1 fusion protein. [1] Mixed lineage leukemia 1 (MLL1) is a histone H3 lysine 4 (H3K4) methyltransferase that interacts with WD repeat domain 5 (WDR5) to regulate cell survival, proliferation and senescence. The role of MLL1 in the pathogenesis of acute kidney injury (AKI) is unclear. In this study, we found that cisplatin-induced upregulation of MLL1, WDR5, and trimethylated H3K4 (H3K4me3) expression in mouse AKI renal tubular cells, along with increased p53 phosphorylation and decreased E-cadherin expression. Administration of the selective MLL1/WDR5 complex inhibitor MM102 improved renal function, reduced renal tubular damage and apoptosis, while inhibiting the expression of MLL1, WDR5, and H3K4me3, dephosphorylating p53, and maintaining E-cadherin expression. In cultured mouse proximal renal tubular cells (RPTCs), cisplatin treatment, MM102 treatment, or transfection with MLL1 or WDR5 siRNA all inhibited apoptosis and p53 phosphorylation while maintaining E-cadherin expression. Inhibition of p53 with Pifithrin-α reduced cisplatin-induced apoptosis but did not affect the expression of MLL1, WDR5, and H3K4me3. Interestingly, silencing E-cadherin counteracts the cytoprotective effects of MM102 but has no effect on p53 phosphorylation. These results suggest that MLL1/WDR5 activates p53, thereby inhibiting E-cadherin and ultimately leading to apoptosis during cisplatin-induced acute kidney injury (AKI). Further studies showed that MM102 effectively inhibits cisplatin-induced DNA damage response (DDR), manifested as dephosphorylation of ataxia-telangiectasia mutant protein (ATM) and ATM and Rad-3-related protein (ATR), dephosphorylation of checkpoint kinases 1 and 2 (Chk1 and Chk2), inhibition of γ-H2AX, and inhibition of cell cycle arrest (manifested as decreased p21 expression in vitro and in vivo and reduced phosphorylation of histone H3 at serine 10). Overall, we found that MLL1 is a novel DNA damage response (DDR) regulator that drives cisplatin-induced renal tubular epithelial cell (RPTC) apoptosis and acute kidney injury (AKI) by regulating the MLL1/WDR5/ATR/ATM-Chk-p53-E-cadherin axis. Targeting the MLL1/WDR5 complex may have the potential to treat AKI. [2] 1. Mechanism of action: MM-102 is a peptide mimic inhibitor of MLL1-WDR5 protein-protein interaction. It binds to the “WIN” site of WDR5, blocking the recruitment of MLL1 to WDR5, thereby disrupting the MLL1 methyltransferase complex. This inhibits H3K4me3 modification on MLL1 target genes (e.g., HOXA9, MEIS1), thereby suppressing oncogenic transcription in leukemia and p53-mediated apoptosis in cisplatin-induced AKI [1][2]
2. Background of leukemia treatment: MLL rearrangement leukemia depends on MLL1-WDR5 interaction to express oncogenes and maintain cell survival. MM-102 targets this dependence and has shown preclinical efficacy in xenograft models, supporting its development for the treatment of MLL rearrangement leukemia [1] 3. AKI therapeutic potential: Cisplatin-induced AKI involves MLL1-mediated E-cadherin inhibition and p53-dependent activation of apoptosis. MM-102 can inhibit MLL1, restore the expression of E-cadherin, and reduce renal tubular cell apoptosis, representing a novel treatment strategy for cisplatin-induced kidney injury [2]. 4. Peptide mimicry design: MM-102 is designed based on the MLL1 “WIN” motif peptide and enhances its binding affinity, stability, and oral bioavailability through structural modifications (e.g., cyclization, hydrophobic substitution)—these are its key advantages over linear peptide inhibitors [1]. |
| Molecular Formula |
C35H49F2N7O4
|
|
|---|---|---|
| Molecular Weight |
669.8
|
|
| Exact Mass |
669.381
|
|
| Elemental Analysis |
C, 62.76; H, 7.37; F, 5.67; N, 14.64; O, 9.55
|
|
| CAS # |
1417329-24-8
|
|
| Related CAS # |
MM-102 TFA;1883545-52-5
|
|
| PubChem CID |
54766613
|
|
| Appearance |
White to off-white solid powder
|
|
| LogP |
6.433
|
|
| Hydrogen Bond Donor Count |
6
|
|
| Hydrogen Bond Acceptor Count |
7
|
|
| Rotatable Bond Count |
16
|
|
| Heavy Atom Count |
48
|
|
| Complexity |
1080
|
|
| Defined Atom Stereocenter Count |
1
|
|
| SMILES |
O=C(C1(NC([C@@H](NC(C(NC(C(C)C)=O)(CC)CC)=O)CCCNC(N)=N)=O)CCCC1)NC(C2=CC=C(F)C=C2)C3=CC=C(F)C=C3
|
|
| InChi Key |
RZKSQRIPRKWVBU-MHZLTWQESA-N
|
|
| InChi Code |
InChI=1S/C35H49F2N7O4/c1-5-34(6-2,43-29(45)22(3)4)31(47)41-27(10-9-21-40-33(38)39)30(46)44-35(19-7-8-20-35)32(48)42-28(23-11-15-25(36)16-12-23)24-13-17-26(37)18-14-24/h11-18,22,27-28H,5-10,19-21H2,1-4H3,(H,41,47)(H,42,48)(H,43,45)(H,44,46)(H4,38,39,40)/t27-/m0/s1
|
|
| Chemical Name |
N-[Bis(4-fluorophenyl)methyl]-1-[[(2S)-5-(diaminomethylideneamino)-2-[[2-ethyl-2-(2-methylpropanoylamino)butanoyl]amino]pentanoyl]amino]cyclopentane-1-carboxamide
|
|
| Synonyms |
|
|
| HS Tariff Code |
2934.99.9001
|
|
| Storage |
Powder -20°C 3 years 4°C 2 years In solvent -80°C 6 months -20°C 1 month Note: Please store this product in a sealed and protected environment (e.g. under nitrogen), avoid exposure to moisture and light. |
|
| Shipping Condition |
Room temperature (This product is stable at ambient temperature for a few days during ordinary shipping and time spent in Customs)
|
| Solubility (In Vitro) |
|
|||
|---|---|---|---|---|
| Solubility (In Vivo) |
Note: Listed below are some common formulations that may be used to formulate products with low water solubility (e.g. < 1 mg/mL), you may test these formulations using a minute amount of products to avoid loss of samples.
Injection Formulations
Injection Formulation 1: DMSO : Tween 80: Saline = 10 : 5 : 85 (i.e. 100 μL DMSO stock solution → 50 μL Tween 80 → 850 μL Saline)(e.g. IP/IV/IM/SC) *Preparation of saline: Dissolve 0.9 g of sodium chloride in 100 mL ddH ₂ O to obtain a clear solution. Injection Formulation 2: DMSO : PEG300 :Tween 80 : Saline = 10 : 40 : 5 : 45 (i.e. 100 μL DMSO → 400 μLPEG300 → 50 μL Tween 80 → 450 μL Saline) Injection Formulation 3: DMSO : Corn oil = 10 : 90 (i.e. 100 μL DMSO → 900 μL Corn oil) Example: Take the Injection Formulation 3 (DMSO : Corn oil = 10 : 90) as an example, if 1 mL of 2.5 mg/mL working solution is to be prepared, you can take 100 μL 25 mg/mL DMSO stock solution and add to 900 μL corn oil, mix well to obtain a clear or suspension solution (2.5 mg/mL, ready for use in animals). View More
Injection Formulation 4: DMSO : 20% SBE-β-CD in saline = 10 : 90 [i.e. 100 μL DMSO → 900 μL (20% SBE-β-CD in saline)] Oral Formulations
Oral Formulation 1: Suspend in 0.5% CMC Na (carboxymethylcellulose sodium) Oral Formulation 2: Suspend in 0.5% Carboxymethyl cellulose Example: Take the Oral Formulation 1 (Suspend in 0.5% CMC Na) as an example, if 100 mL of 2.5 mg/mL working solution is to be prepared, you can first prepare 0.5% CMC Na solution by measuring 0.5 g CMC Na and dissolve it in 100 mL ddH2O to obtain a clear solution; then add 250 mg of the product to 100 mL 0.5% CMC Na solution, to make the suspension solution (2.5 mg/mL, ready for use in animals). View More
Oral Formulation 3: Dissolved in PEG400 (Please use freshly prepared in vivo formulations for optimal results.) |
| Preparing Stock Solutions | 1 mg | 5 mg | 10 mg | |
| 1 mM | 1.4930 mL | 7.4649 mL | 14.9298 mL | |
| 5 mM | 0.2986 mL | 1.4930 mL | 2.9860 mL | |
| 10 mM | 0.1493 mL | 0.7465 mL | 1.4930 mL |
*Note: Please select an appropriate solvent for the preparation of stock solution based on your experiment needs. For most products, DMSO can be used for preparing stock solutions (e.g. 5 mM, 10 mM, or 20 mM concentration); some products with high aqueous solubility may be dissolved in water directly. Solubility information is available at the above Solubility Data section. Once the stock solution is prepared, aliquot it to routine usage volumes and store at -20°C or -80°C. Avoid repeated freeze and thaw cycles.
Calculation results
Working concentration: mg/mL;
Method for preparing DMSO stock solution: mg drug pre-dissolved in μL DMSO (stock solution concentration mg/mL). Please contact us first if the concentration exceeds the DMSO solubility of the batch of drug.
Method for preparing in vivo formulation::Take μL DMSO stock solution, next add μL PEG300, mix and clarify, next addμL Tween 80, mix and clarify, next add μL ddH2O,mix and clarify.
(1) Please be sure that the solution is clear before the addition of next solvent. Dissolution methods like vortex, ultrasound or warming and heat may be used to aid dissolving.
(2) Be sure to add the solvent(s) in order.
| NCT Number | Recruitment | interventions | Conditions | Sponsor/Collaborators | Start Date | Phases |
| NCT01997840 | Active, not recruiting | Drug: ACY-1215 (Ricolinostat) in combination with pomalidomide and dexamethasone |
Multiple Myeloma | Celgene | March 1, 2014 | Phase 1 Phase 2 |
| NCT02426723 | Completed | Drug: Phase 1a: CWP232291
Drug: Phase 1b: CWP232291, Lenalidomide, Dexamethasone |
Multiple Myeloma |
JW Pharmaceutical | October 19, 2015 | Phase 1 |
| NCT00985959 | Completed Has Results | Drug: JNJ-26866138 0.7 mg/m2 Drug: JNJ-26866138 1.0 mg/m2 |
Multiple Myeloma |
Janssen Pharmaceutical K.K. | July 2008 | Phase 1 |
 |
|---|