
| Size | Price | Stock | Qty |
|---|---|---|---|
| 250mg |
|
||
| 500mg |
|
||
| 1g |
|
||
| 2g |
|
||
| Other Sizes |
Purity: =99.85%
Emodin (HSDB-7093; NSC-408120; NSC-622947; Emodol; Frangula emodin), a naturally occuring anthraquinone analog found in rubarb, buckthorn, and Japanese knotweed, is a potent and broad-spectrum anticancer agent that also has anti-inflammatory and neuroprotective effects. It inhibits casein kinase II (CKII) activity with IC50 of 2 μM.
| Targets |
Natural anthraquinone; casein kinase-2 (CK2); SARS-CoV; CK2α; 11β-HSD1
SARS Coronavirus (SARS-CoV) Spike protein (S protein)-Angiotensin-converting enzyme 2 (ACE2) interaction: Emodin blocks the binding between SARS-CoV S protein and host cell ACE2, with no enzyme activity-related IC50/Ki values (targets protein-protein interaction rather than enzyme) [1] - Casein Kinase 2 (CK2): Emodin is a selective inhibitor of human recombinant CK2, with an IC50 of 2.3 μM (measured by inhibiting CK2-mediated phosphorylation of GST-Myb substrate) [2] - 11β-Hydroxysteroid dehydrogenase type 1 (11β-HSD1): Emodin inhibits human recombinant 11β-HSD1 activity, with an IC50 of 1.7 μM (measured by suppressing the conversion of cortisone to cortisol) [3] |
|---|---|
| ln Vitro |
Emodin (10–400 μM) has an IC50 value of 200 μM and inhibits S protein binding to ACE2 in a dose-dependent manner[1]. In a dose-dependent manner, emodin (5-50 μM) decreases the infectivity of S-protein pseudotyped retroviruses. The SARS-CoV S protein cannot attach to Vero E6 cells when imodin is present [1]. Emodin inhibits casein kinase 2 (CK2) at ATP concentrations of 50 μM, 30.0 μM, and 7.1 μM for CK2α wild type, Ile174Ala mutant, and His160Ala mutant, respectively, with an IC50 of 5.9. At ATP concentration, the IC50 values for the CK2α wild type and the Val66Ala mutant are 1.40 and 38.00 μM, respectively [2]. Emodin is more than 5000 times more selective for the human and mouse 11β-hydroxysteroid dehydrogenase type 2 (11β-HSD2) enzyme than type 2 isoenzymes, as seen by its modest inhibitory action against both mouse and human 11β-HSD2 (IC50 greater than 1 mM) [3].
Emodin, an anthraquinone compound derived from genus Rheum and Polygonum, significantly blocked the S protein and ACE2 interaction in a dose-dependent manner. It also inhibited the infectivity of S protein-pseudotyped retrovirus to Vero E6 cells. These findings suggested that emodin may be considered as a potential lead therapeutic agent in the treatment of SARS. [1] Emodin is a potent and selective 11beta-HSD1 inhibitor with the IC(50) of 186 and 86 nM for human and mouse 11beta-HSD1, respectively [3]. Anti-SARS-CoV activity: - S protein-ACE2 binding inhibition: ELISA-based binding assay showed that Emodin (50 μM) reduced the binding of recombinant S protein to ACE2-coated plates by 80%; surface plasmon resonance (SPR) analysis confirmed a dose-dependent decrease in binding affinity (Kd increased from 0.2 nM to 2.5 nM at 30 μM Emodin) [1] - Viral replication inhibition: In Vero cells infected with SARS-CoV (MOI=0.1), Emodin (10 μM) reduced viral titer by 100-fold (from 1×10⁶ TCID50/mL to 1×10⁴ TCID50/mL) after 48 h incubation. Cell viability (MTT assay) showed CC50=85 μM, giving a selectivity index (SI=CC50/EC50) of 8.5 [1] - CK2 inhibition and anti-proliferation: - Recombinant CK2 activity: Emodin (0.5-10 μM) dose-dependently inhibited human recombinant CK2, with 2.3 μM IC50. Radioautography showed 10 μM Emodin reduced CK2-mediated phosphorylation of GST-Myb by 70% [2] - HeLa cell proliferation: Emodin (5-50 μM) suppressed HeLa cell growth, with IC50=15 μM (MTT assay, 72 h). Western blot revealed downregulated phosphorylated Myb (p-Myb, a CK2 substrate) and cyclin D1 (cell cycle marker) at 10 μM [2] - 11β-HSD1 inhibition and metabolic regulation: - Hepatic 11β-HSD1 activity: In mouse liver microsomes, Emodin (0.5-5 μM) inhibited 11β-HSD1-mediated cortisone-to-cortisol conversion by 30-80%; 1.7 μM achieved 50% inhibition [3] - Adipocyte insulin sensitivity: In differentiated 3T3-L1 adipocytes, Emodin (10-20 μM) increased insulin-stimulated AKT phosphorylation (Ser473) by 1.8-2.5-fold (Western blot) and reduced high glucose-induced reactive oxygen species (ROS) by 40% (DCFH-DA staining, flow cytometry) [3] |
| ln Vivo |
In normal C57BL/6J male mice, emodin (single oral treatment of 100 or 200 mg/kg) suppresses 11β-HSD1 activity [3]. Emodin (100 mg/kg; oral; bid) lowers blood glucose, liver PEPCK, and glucose-6-phosphatase mRNA in diet-induced obesity (DIO) rats, improving insulin sensitivity and lipid metabolism [3].
Single oral administration of emodin inhibited 11beta-HSD1 activity of liver and fat significantly in mice. Emodin reversed prednisone-induced insulin resistance in mice, whereas it did not affect dexamethasone-induced insulin resistance, which confirmed its inhibitory effect on 11beta-HSD1 in vivo. In DIO mice, oral administration of emodin improved insulin sensitivity and lipid metabolism, and lowered blood glucose and hepatic PEPCK, and glucose-6-phosphatase mRNA. Conclusions and implications: This study demonstrated a new role for emodin as a potent and selective inhibitor of 11beta-HSD1 and its beneficial effects on metabolic disorders in DIO mice. This highlights the potential value of analogues of emodin as a new class of compounds for the treatment of metabolic syndrome or type 2 diabetes.[3] Diet-induced obese (DIO) mouse model (metabolic disorder): - Animals and treatment: Male C57BL/6 mice (6 weeks old) were fed a high-fat diet (HFD, 60% fat) for 12 weeks to induce obesity, then randomized into 3 groups (n=8/group): HFD + vehicle (0.5% CMC-Na, p.o.), HFD + Emodin 10 mg/kg/day (p.o.), HFD + Emodin 30 mg/kg/day (p.o.). Normal diet (ND) group served as control. Treatment duration was 4 weeks [3] - Metabolic outcomes: 30 mg/kg Emodin significantly improved metabolic parameters vs. HFD vehicle: fasting blood glucose (13.2 ± 1.5 mmol/L vs. 17.6 ± 1.8 mmol/L, -25%), serum insulin (28 ± 4 μU/mL vs. 40 ± 5 μU/mL, -30%), total cholesterol (4.8 ± 0.4 mmol/L vs. 6.5 ± 0.6 mmol/L, -26%), triglycerides (1.8 ± 0.2 mmol/L vs. 2.8 ± 0.3 mmol/L, -36%) [3] - Tissue effects: Hepatic 11β-HSD1 activity was reduced by 50% in 30 mg/kg group; white adipose tissue (WAT) mRNA levels of pro-inflammatory cytokines (TNF-α, IL-6) decreased by 45% and 50%, respectively (qPCR). No significant weight change was observed (HFD vehicle: 28.5 ± 1.2 g vs. 30 mg/kg Emodin: 27.8 ± 1.0 g) [3] |
| Enzyme Assay |
For the competition assay, biotinylated S protein was mixed with various amounts of extracts and incubated at 37 °C with shaking. After a 2-h incubation, the mixture was added to wells, which were coated with ACE2, and incubated at 37 °C for 1 h. Following three washes, peroxidase-conjugated avidin and chromatic substrate were sequentially added. The absorbance was read at 405 nm in an ELISA plate reader. The percent inhibition was calculated by [1 − (OD value of mixture containing extract and S protein/OD value of mixture containing S protein only)] × 100. The IC50 value was determined as the quantity of compound required to inhibit the interaction between S protein and ACE2 at 50%.[1]
Phosphorylation assay[2] Phosphorylation assays were carried out in the presence of increasing amounts of each inhibitor tested in a final volume of 25 μL containing 50 mM Tris-HCl, pH 7.5, 100 mM NaCl, 12 mM MgCl2, 100 μM synthetic peptide substrate RRRADDSDDDDD, and 0.02 mM [γ-33P]ATP (500–1000 cpm/pmol), unless otherwise indicated, and incubated for 10 min at 37 °C. These conditions are suited for reaching maximal velocity. Assays were stopped by addition of 5 μL of 0.5 M orthophosphoric acid before spotting aliquots onto phosphocellulose filters. Filters were washed in 75 mM phosphoric acid (5–10 mL/each) four times, then once in methanol and dried before counting.[2] Measurement of 11β-HSD1 and -HSD2 activity in vitro[3] The SPA was used to screen for inhibitors of 11β-HSDs (Mundt et al., 2005), with the microsome fractions prepared from the HEK-293 cells stably transfected with either human or mouse 11β-HSD1 or 11β-HSD2 as the enzyme source. Briefly, different concentrations of compound were added to 96-well microtitre plates, followed by the addition of 80 µL of 50 mM HEPES buffer, pH 7.4 containing 25 nM [1,2-(n)3H]-cortisone and 1.25 mM NADPH (for 11β-HSD1 assay) or 12.5 nM [1,2,6,7-(n)3H]-cortisol and 0.625 mM NAD+(for 11β-HSD2 assay). Reactions were initiated by the addition of 11β-HSD1 or 11β-HSD2, enzyme preparation as microsome fractions from HEK293 cells in a final concentration of 80 µg·mL−1 for 11β-HSD1, and 160 µg·mL−1 for 11β-HSD2, respectively. After a 60 min incubation at 37°C, the reaction was stopped by the addition of 35 µL of 10 mg·mL−1 protein A-coated yttrium silicate beads suspended in SuperBlock Blocking Buffer with 3 µg·mL−1 of murine monoclonal cortisol antibody and 314 µM glycyrrhetinic acid. The plates were incubated under plastic film on an orbital shaker for 120 min at room temperature before counting. The amount of [3H]-cortisol generated in 11β-HSD1 enzyme reaction or remaining from the 11β-HSD2 enzyme reaction was captured by the beads and determined in a microplate liquid scintillation counter. The % inhibition was calculated relative to a non-inhibited control. Data were obtained from at least three independent experiments. IC50 values were calculated from concentration–response curves by a non-linear regression analysis using Prism Version 4. SARS-CoV S protein-ACE2 binding assay (SPR): 1. Sensor chip preparation: Immobilize recombinant human ACE2 (10 μg/mL) onto a CM5 sensor chip via amine coupling, with a surface density of ~1500 RU. 2. Binding reaction: Inject a mixture of recombinant SARS-CoV S protein (0.1 μM) and Emodin (0, 10, 30, 50, 100 μM) into the SPR system at 30 μL/min, 25℃. 3. Signal detection: Record the association (60 s) and dissociation (120 s) phases. Calculate binding affinity (Kd) using BIAevaluation software. Binding inhibition rate = [(RUcontrol - RUtreatment)/RUcontrol] × 100% [1] - CK2 activity assay: 1. Reaction system: Prepare 50 μL mixtures containing 50 mM Tris-HCl (pH 7.5), 10 mM MgCl₂, 0.1 mM ATP (including 0.5 μCi [γ-³²P]-ATP), 2 μg GST-Myb substrate, 0.5 μg recombinant human CK2, and Emodin (0, 0.5, 1, 2.3, 5, 10 μM). 2. Incubation and termination: Incubate at 37℃ for 30 min. Add 12 μL 5×SDS-PAGE loading buffer (containing β-mercaptoethanol) to terminate the reaction. 3. Detection and quantification: Separate proteins by 12% SDS-PAGE, dry the gel, and expose to X-ray film for radioautography. Quantify the intensity of phosphorylated GST-Myb bands using ImageJ software. CK2 activity = (band intensity of treatment/band intensity of control) × 100% [2] - 11β-HSD1 activity assay: 1. Reaction system: Prepare 100 μL mixtures containing 100 mM Tris-HCl (pH 7.4), 200 μM NADPH, 10 μM cortisone (substrate), 50 μg mouse liver microsomes (or 2 μg recombinant human 11β-HSD1), and Emodin (0, 0.5, 1, 1.7, 3, 5 μM). 2. Incubation and termination: Incubate at 37℃ for 60 min. Add 200 μL ice-cold methanol to terminate the reaction, vortex for 30 s, and centrifuge at 12,000 × g for 10 min at 4℃. 3. Cortisol detection: Collect the supernatant, filter through a 0.22 μm membrane, and analyze by HPLC (C18 column, 4.6×250 mm). Mobile phase: methanol:water (60:40, v/v), flow rate 1 mL/min, detection wavelength 240 nm. Calculate cortisol concentration using a standard curve. 11β-HSD1 activity = nmol cortisol formed per mg protein per hour [3] |
| Cell Assay |
Cell Viability Assay[1]
Cell Types: Vero E6 cells transfected with the plasmid encoding ACE2 Tested Concentrations: 0, 5, 25, 50 μM Incubation Duration: 24 hrs (hours) Experimental Results: Vero cells treated with 50 μM remained 82.4±3.8% viability, the anti -SARS-CoV activity was not due to toxicity. Biotinylated ELISA [1] The biotinylated ELISA was performed as described previously (Ho et al., 2006). Briefly, microtiter plates were coated at 4 °C overnight with 50 μl of 0.2 ng/μl ACE2, rinsed with 200 μl washing buffer (0.5% Tween 20 in phosphate-buffered saline (PBS) (137 mM NaCl, 1.4 mM KH2PO4, 4.3 mM Na2HPO4, 2.7 mM KCl, pH 7.2)), and blocked with 200 μl blocking buffer (5% bovine serum albumin (BSA) in washing buffer) by incubating at 37 °C for 30 min. The absorbed protein in each well was challenged with 50 μl of 1 ng/μl biotinylated S protein and incubated at 37 °C for 1 h. After three washes with washing buffer, 50 μl diluted peroxidase-conjugated avidin was added to each well and incubated at 37 °C for 1 h. Following three washes, 50 μl chromogenic substrate, 2,2′-azino-bis(3-ethylbenzothiazoline-6-sulfonic acid), was added to each well and incubated at 37 °C for 15 min. The absorbance was read at 405 nm in an ELISA plate reader. For the competition assay, biotinylated S protein was mixed with various amounts of extracts and incubated at 37 °C with shaking. After a 2-h incubation, the mixture was added to wells, which were coated with ACE2, and incubated at 37 °C for 1 h. Following three washes, peroxidase-conjugated avidin and chromatic substrate were sequentially added. The absorbance was read at 405 nm in an ELISA plate reader. The percent inhibition was calculated by [1 − (OD value of mixture containing extract and S protein/OD value of mixture containing S protein only)] × 100. The IC50 value was determined as the quantity of compound required to inhibit the interaction between S protein and ACE2 at 50%. Immunofluorescence assay (IFA) [1] Vero E6 cells (104 cells) were seeded in 24-well plates containing glass coverslips and incubated at 37 °C for 1 day. The coverslips were then rinsed with PBS, fixed with 3.7% PBS-buffered formaldehyde at room temperature for 30 min, and blocked with 1% BSA at 37 °C for 1 h. After four washes with PBS, biotin-labeled S protein was added to each coverslip and incubated at 4 °C overnight. Following four washes with PBS, diluted fluorescence-conjugated streptavidin was added and incubated at 37 °C for 90 min in the dark. The coverslips were then washed four times with PBS, placed onto glass slides, mounted with fluoromount G, and observed under a confocal microscope. SARS-CoV infection assay in Vero cells: 1. Cell seeding: Seed Vero cells into 96-well plates at 1×10⁴ cells/well, incubate in DMEM (10% FBS) at 37℃, 5% CO₂ for 24 h until 80% confluence. 2. Drug pretreatment and viral infection: Replace medium with fresh DMEM containing Emodin (0, 5, 10, 20, 50, 100 μM) and incubate for 1 h. Then add SARS-CoV (MOI=0.1) to each well, incubate for 2 h, and replace with DMEM (2% FBS) containing the same Emodin concentration. 3. Viral titer and cell viability: After 48 h, observe cytopathic effect (CPE) under a microscope. Determine viral titer by TCID50 method. For viability, add 20 μL MTT (5 mg/mL) to each well, incubate for 4 h, dissolve formazan with 150 μL DMSO, and measure OD570 nm [1] - HeLa cell proliferation and CK2 substrate assay: 1. Proliferation assay: Seed HeLa cells into 96-well plates at 5×10³ cells/well, incubate for 24 h. Add Emodin (0, 5, 10, 15, 20, 50 μM), incubate for 72 h, and perform MTT assay as above to calculate IC50. 2. Western blot for p-Myb: Seed HeLa cells into 6-well plates at 2×10⁵ cells/well, incubate for 24 h. Treat with Emodin (0, 5, 10, 20 μM) for 48 h, lyse cells with RIPA buffer (containing phosphatase inhibitors). Separate proteins by SDS-PAGE, transfer to PVDF membrane, incubate with anti-p-Myb (1:1000) and anti-GAPDH (1:5000) antibodies, then with HRP-conjugated secondary antibody. Detect signals by ECL and quantify with ImageJ [2] - 3T3-L1 adipocyte insulin signaling and ROS assay: 1. Insulin signaling: Differentiate 3T3-L1 preadipocytes into adipocytes (using IBMX, dexamethasone, insulin). Treat with Emodin (0, 10, 20 μM) for 24 h, stimulate with insulin (100 nM) for 15 min, lyse cells, and perform Western blot for p-AKT (Ser473) and total AKT. 2. ROS detection: Load adipocytes with 10 μM DCFH-DA for 30 min at 37℃, treat with high glucose (25 mM) + Emodin (10 μM) for 2 h. Measure fluorescence intensity (excitation 488 nm, emission 525 nm) using a flow cytometer. ROS level = (fluorescence of treatment/fluorescence of control) × 100% [3] |
| Animal Protocol |
Animal/Disease Models: C57BL/6J male mice[3]
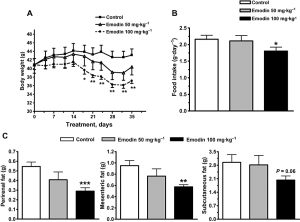
\nDoses: 100 or 200 mg/kg \nRoute of Administration: Acute administered po; Two hrs (hours) later, the mice were killed by cervical dislocation, \nExperimental Results: Dramatically inhibited liver 11β-HSD1 enzymatic activity by 17.6 and 31.3% and mesenteric fat 11β-HSD1 enzymatic activity by 21.5 and 46.7% at 100 or 200 mg/kg, respectively. \n \nAnimal/Disease Models: DIO mice (C57BL/6J male mice were fed a formulated research diet)[3] \nDoses: 100 mg /kg \nRoute of Administration: po (oral gavage); twice per day; for 35 days \nExperimental Results: decreased fasting glucose concentrations to 77.2% of the vehicle control mice after 7 days of treatment, and these remained Dramatically lower throughout the treatment period. demonstrated a significant reduction in blood glucose levels at all time-points following oral glucose challenge after 24 days of treatment. Evoked a Dramatically greater reduction in blood glucose values 40 and 90 min after insulin injection after 28 days of treatment. The serum insulin level was also significant. \n Acute administration in normal mice [3] \nTo evaluate the activity of acute administration of Emodin, C57 BL/6J mice deprived of food overnight were administered emodin (100 or 200 mg·kg−1) or vehicle (0.5% carboxymethylcellulose; CMC) p.o. Two hours later, animals were killed by cervical dislocation, and the liver and mesenteric fat were isolated immediately, washed in ice-cold PBS, frozen in liquid nitrogen and stored at −80°C. The liver and mesenteric fat were homogenized (0.1 g·mL−1) in cold homogenization buffer (20 mM Na2HPO4, 5% glycerol, 1 mM EDTA, pH 7.0), and 10 µg of liver homogenates or 30 µg mesenteric fat homogenates was used to analyse the 11β-HSD1 activity by SPA, as previously described. \n Effect of Emodin on prednisone- or dexamethasone-induced insulin-resistant mice [3] \nMale C57BL/6J mice (8 weeks old) were randomly assigned to six groups based on body weight. The experimental groups and respective treatment were as follows: (i) control – 0.5% CMC; (ii) prednisone acetate – 100 mg·kg−1; (iii) prednisone acetate (100 mg·kg−1) plus emodin (100 mg·kg−1); (iv) prednisone acetate (100 mg·kg−1) plus Emodin (200 mg·kg−1); (v) dexamethasone (4 mg·kg−1); and (vi) dexamethasone (4 mg·kg−1) plus emodin (200 mg·kg−1). Prednisone or dexamethasone was administered by oral gavage twice daily to induce a state of glucocorticoid excess and insulin resistance in mice. Emodin was administered orally twice daily 1 day before, and then at the same time as prednisone or dexamethasone. After 14 days of treatment, insulin tolerance was determined in mice deprived of food overnight (0.5 U·kg−1 insulin administered by an i.p. injection) to investigate the effect of emodin on prednisone- or dexamethasone-induced insulin resistance. \n Effect of Emodin in DIO mice [3] \nC57BL/6J male mice (3–4 weeks) were fed a formulated research diet containing 60% of the calories from fat for 12 weeks before, and throughout the duration of the experiment. DIO mice were assigned to three groups and subjected to gavage treatment twice per day with vehicle (0.5% CMC), Emodin 50 or 100 mg·kg−1, respectively, for 35 days. Fasting blood glucose values and initial body weights were comparable between groups. The blood glucose levels were measured via blood drops obtained by clipping the tail of the mice using a ONE TOUCH BASIC plus Glucose Monitor, unless otherwise specified. The food intake and body weight of the animals were recorded every 3 days. Glucose tolerance test was determined in mice deprived of food for 5 h (2 g·kg−1 glucose administered by gavage) at day 24 of the treatment. The blood samples were collected via the retro-orbital sinus, and the serum glucose and insulin concentrations were measured with an enzymatic colorimetric method and insulin elisa kit, respectively. An insulin tolerance test was performed in the 5 h-fasted mice (0.5 U·kg−1 insulin administered by an i.p. injection) at day 28 of the treatment. On the last day of treatment, 5 h-fasted mice were anaesthetized with an i.p. injection of sodium pentobarbital (40 mg·kg−1). Serum was collected for determination of insulin, triacylglycerol, cholesterols and non-esterified free fatty acid (NEFA) concentration. The liver and different fat pads including epididymal fat, mesenteric fat, perirenal fat and subcutaneous fat were dissected, weighed, immediately frozen in liquid nitrogen and stored at −80°C. \nDIO Mouse Model for Metabolic Disorder: \n1. Animal selection and acclimation: Male C57BL/6 mice (6 weeks old, 18-20 g) were housed under SPF conditions (12 h light/dark cycle, 22±2℃), free access to food and water. Acclimate for 1 week before the experiment. \n2. Obesity induction: Mice were divided into ND group (normal chow, 10% fat) and HFD group (60% fat diet) for 12 weeks. Mice with body weight ≥25 g (HFD group) were selected for subsequent treatment. \n3. Grouping and drug administration: Selected HFD mice were randomized into 2 subgroups (n=8/group): \n- HFD vehicle: Oral gavage of 0.5% carboxymethylcellulose sodium (CMC-Na) once daily. \n- HFD + Emodin: Oral gavage of Emodin (10 or 30 mg/kg/day, dissolved in 0.5% CMC-Na, sonicated to dissolve). \nND group (n=8) received oral gavage of 0.5% CMC-Na. Treatment duration: 4 weeks. \n4. Sample collection and detection: \n- Weekly measurements: Record body weight and food intake every week. \n- Metabolic parameters: At the end of treatment, mice were fasted for 12 h, blood was collected from orbital sinus. Serum glucose (glucose oxidase method), insulin (ELISA), total cholesterol (TC), and triglycerides (TG) (enzymatic method) were measured. \n- Tissue analysis: Mice were euthanized by cervical dislocation. Liver was collected to measure 11β-HSD1 activity; WAT was collected to extract RNA for qPCR (TNF-α, IL-6); liver and kidney were fixed in 4% paraformaldehyde for HE staining [3] |
| ADME/Pharmacokinetics |
Absorption, Distribution and Excretion
This study investigated the absorption, excretion, tissue distribution, and metabolism of the anthraquinone compound [14C]emodin in rats following a single oral administration (approximately 50 mg/kg). Urinary excretion was 18 (±5)% of the administered dose within 24 hours and 22 (±6)% within 72 hours. In urinary samples from 0–72 hours, the main metabolites were free anthraquinone compounds (emodin and rhein, accounting for 16% of the administered dose); 3% were bound compounds, and 3% were unextractable radioactive substances. Fecal excretion was 48 ± 11% within 24 hours and 68 ± 8% within 120 hours, primarily in the form of free anthraquinone compounds. In two cannulated rats, bile excretion peaked at approximately 6 hours and reached 49% of the dose within 15 hours; 70% of the bile radioactivity was in the form of bound emodin. The radioactivity levels in most organs decreased significantly within 3 to 5 days. However, in the kidneys, the radioactivity of 14C remained equivalent to 4.33 ppm of emodin after 5 days. The radioactivity of 14C in the mesentery and adipose tissue gradually increased over 72 to 120 hours. Metabolism/Metabolites…The metabolism of emodin (1,3,8-trihydroxy-6-methylanthraquinone) was investigated…The formation of two emodin metabolites—ω-hydroxyemodin and 2-hydroxyemodin—was observed in rat liver microsomes. The rate of ω-hydroxyemodin production in rat liver microsomes pretreated with different cytochrome P450 enzyme inducers did not differ. Therefore, the production of ω-hydroxyemodin appears to be catalyzed by several cytochrome P450 enzymes at a low rate. Pretreatment with 3-methylcholanthrene in rat liver microsomes increased the production of 2-hydroxyemodin, which was inhibited by α-naphthoflavone, anti-rat cytochrome P450 1A1/2 antibody, and (to a lesser extent) anti-rat cytochrome P450 1A1 antibody. These data suggest that cytochrome P450 1A2 is involved in the production of this metabolite. However, other cytochrome P450 enzymes also appear to catalyze this reaction. The anthraquinone compound emodin (1,8-dihydroxy-3-methylanthraquinone) is converted to aloe-emodin (1,8-dihydroxy-3-hydroxymethylanthraquinone) via cytochrome P450-dependent oxidation, which is the major metabolite. Liver microsomes from different animals can convert emodin (1,3,8-trihydroxy-6-methylanthraquinone) into an unidentified anthraquinone compound, as well as 2-hydroxyemodin, 4-hydroxyemodin, and 7-hydroxyemodin. ...The major metabolite was identified as ω-hydroxyemodin (1,3,8-trihydroxy-6-hydroxymethylanthraquinone). Among seven animal species, guinea pigs and rats exhibited the highest activity of ω-hydroxyemodin production via liver microsomes, followed by mice and rabbits. Pretreatment of animals with phenobarbital accelerated the microsomal conversion of emodin to ω-hydroxyemodin, while SKF 525A inhibited this activity. It is speculated that the microsomal hydroxylation of the emodin methyl residues and anthraquinone nucleus is specifically catalyzed by multiple forms of cytochrome P-450 regions. ...Emodin is biotransformed via microsomal enzymes into at least five quinone metabolites, one of which, 2-hydroxyemodin (1,2,3,8-tetrahydroxy-6-methylanthraquinone), has been identified as having direct mutagenic effects on the test strains. The activity of the other four quinone metabolites was negative or much lower than that of the active ingredient. Known metabolites of emodin include emodin 3-hydroxyglucuronide. Emodin is biotransformed into active hydroxyemodin, such as ω-hydroxyemodin and 2-hydroxyemodin, by microsomal cytochrome P450 enzymes. Emodin glycosides enter the large intestine unabsorbed and are metabolized by intestinal flora into active aglycones. (A3043, A3046) |
| Toxicity/Toxicokinetics |
Toxicity Summary
Emodin exhibits moderate cytotoxicity, inhibiting the growth of multiple cell types by interfering with the cell cycle, likely through stimulation of p53 and p21 expression. It may also exert this effect by causing DNA strand breaks and/or non-covalently binding to DNA and inhibiting the catalytic activity of topoisomerase II. It may also induce apoptosis by inhibiting the electron transport chain and generating reactive oxygen species. Emodin is a potent inhibitor of tyrosine protein kinase Lck and other tyrosine kinase receptors, which may be one reason for its inhibitory effect on cell growth. It may exert a chemopreventive effect by activating DNA repair mechanisms. Emodin can also inhibit metastasis by interfering with the activity of matrix metalloproteinases, through mechanisms including direct interference or inhibition of focal adhesion kinase, mitogen-activated protein kinase, and RAC-α serine/threonine protein kinase activation, as well as partial inhibition of the transcriptional activity of transcription factors AP-1 and nuclear factor NF-κB. Emodin exerts its laxative effect by acting directly or indirectly on colonic epithelial cells. This activates the underlying smooth muscle cells, leading to muscle contraction. Possible mechanisms of this effect include enhancing motilin secretion, activating muscarinic receptors by triggering acetylcholine release, stimulating the protein kinase C-α pathway to improve calcium sensitivity, inhibiting somatostatin secretion, increasing fluid electrolyte accumulation in the distal ileum and colon, and inhibiting the activity of Na+/K+-ATPase and/or potassium channels. The anti-inflammatory effect of emodin stems from its specific inhibition of the transcription factor NF-κB. It can also regulate angiogenesis by inhibiting casein kinase II and nitric oxide synthase and exhibits a strong affinity for estrogen receptor binding. Emodin can induce the microsomal enzyme cytochrome P-450 1A1, thereby sustainably activating its own metabolism. (A3043, A3044, A3045, A3046, A3047, A3048, A3049, A3050, A3051, A3052, A3053, A3054) Toxicity Overview Emodin exhibits moderate cytotoxicity, inhibiting the growth of various cell types by interfering with the cell cycle, possibly through stimulating the expression of p53 and p21. Furthermore, it may also exert this effect by causing DNA strand breaks and/or non-covalently binding to DNA and inhibiting the catalytic activity of topoisomerase II. It may also induce apoptosis by inhibiting the electron transport chain and generating reactive oxygen species. Emodin is a potent inhibitor of tyrosine protein kinase Lck and other tyrosine kinase receptors, which may be one reason for its inhibitory effect on cell growth. Emodin may exert its chemopreventive effect by activating DNA repair mechanisms. It can also inhibit metastasis by interfering with the activity of matrix metalloproteinases. Its mechanisms of action include direct or indirect inhibition of the activation of focal adhesion kinase, mitogen-activated protein kinase, and RAC-α serine/threonine protein kinase, as well as partial inhibition of the transcriptional activity of transcription factors AP-1 and nuclear factor NF-κB. Emodin exerts its laxative effect by acting directly or indirectly on colonic epithelial cells. This activates the underlying smooth muscle cells, leading to muscle contraction. Its mechanisms of action may include enhancing motilin secretion, activating muscarinic receptors by triggering acetylcholine release, stimulating the C-α protein kinase pathway to increase calcium sensitivity, inhibiting somatostatin secretion, increasing fluid electrolyte accumulation in the distal ileum and colon, and inhibiting the activity of Na+/K+-ATPase and/or potassium channels. The anti-inflammatory effect of emodin stems from its specific inhibition of the transcription factor NF-κB. It can also regulate angiogenesis by inhibiting casein kinase II and nitric oxide synthase and exhibits a strong affinity for estrogen receptor binding. Emodin can induce the expression of the microsomal enzyme cytochrome P-450 1A1, thereby continuously activating its own metabolism. Non-human toxicity values LD50: 35 mg/kg (intraperitoneal injection, mouse) (L135) Toxicity data LD50: 35 mg/kg (intraperitoneal injection, mouse) (L135) Non-human toxicity values Mouse intraperitoneal injection LD50 35 mg/kg In vitro toxicity: -Vero cells:Emodin showed low cytotoxicity, CC50=85 μM; cell viability remained above 80% at concentrations ≤50 μM (MTT method, 48 hours)[1] -HeLa cells: CC50=45 μM; <20 μM emodin did not induce significant apoptosis (Annexin V-FITC/PI staining: apoptosis rate <5%)[2] -Primary mouse hepatocytes: Emodin (0-30 μM) showed no significant cytotoxicity, LDH release <10%, and cell viability >85% (MTT method) [3] - In vivo toxicity: - DIO mice: emodin (30 mg/kg/day, 4 weeks) did not cause changes in body weight (compared to HFD control group: 27.8±1.0 g vs. 28.5±1.2 g), serum liver function indicators (ALT: 25±3 U/L vs. 26±4 U/L; AST: 40±5 U/L vs. 42±6 U/L) or kidney function indicators (BUN: 14±2 mg/dL vs. 15±2 mg/dL; creatinine: 0.7±0.1 mg/dL vs. 0.8±0.1 mg/dL). Liver and kidney HE staining showed no necrosis, inflammation, or renal tubular damage [3] |
| References |
|
| Additional Infomation |
Emodin is an orange needle-like or powdery substance. (NTP, 1992)
Emodin is a trihydroxyanthraquinone with the structure 9,10-anthraquinone, where hydroxyl groups are substituted at positions 1, 3, and 8, and a methyl group is substituted at position 6. It is found in the roots and bark of various plants (especially rhubarb and rhamnos), as well as in molds and lichens. It is an active ingredient in many traditional Chinese medicines. Emodin has various effects, including acting as a tyrosine kinase inhibitor, antitumor drug, laxative, and plant metabolite. Its function is similar to that of emodin anthrone. It is the conjugate acid of emodin (1-). Emodin has been studied for the treatment of polycystic kidney disease. Emodin has been reported to exist in Hamigera avellanea, Setophoma terrestris, and other organisms with relevant data. Emodin is found in Sophora sagrada. Emodin is a laxative resin extracted from rhubarb, Polygonum cuspidatum, Hippophae rhamnoides, and Fallopia japonica. The term may also refer to any of a series of compounds isomers of rhubarb emodin. (Wikipedia) Emodin has been shown to possess anti-inflammatory, signal transduction, antibacterial, muscle growth, and anti-angiogenic properties (A3049, A7853, A7854, A7855, A7857). Emodin is a laxative anthraquinone found in a variety of plants, particularly Rhamnus purshiana. It was once used as a laxative but is now primarily used as a tool in toxicology studies. See also: Rhamnus bark (part); Rhamnus renotria root (part). Mechanism of Action This study investigated the inhibitory effects of the anthraquinone fungal toxins emodin and azuril on mouse leukemia L1210 cell culture, rat liver mitochondrial oxidative phosphorylation, and rat brain microsomal Na+,K+ activated ATPase activity to explore the differences in their toxic mechanisms of action. Azuril showed stronger growth inhibition on L1210 cell culture than emodin. Emodin had a stronger uncoupling effect on mitochondrial respiration than azuril. Azuril inhibited rat brain microsomal Na+,K+ activated ATPase activity, while emodin did not have this inhibitory effect. …Emodin can induce apoptosis in human hepatocellular carcinoma (HCC) lines Mahlavu, PLC/PRF/5, and HepG2. The addition of emodin to these three cell lines inhibited cell growth in a time- and dose-dependent manner. Emodin generates reactive oxygen species (ROS) in these cells, leading to a decrease in intracellular mitochondrial transmembrane potential (Δ), which in turn activates caspase-9 and caspase-3, ultimately resulting in DNA fragmentation and apoptosis. Emodin inhibits the activity of TPK and CK2, as well as the degradation of I-κB. …Emodin-induced apoptosis in CH27 cells does not involve the regulation of endogenous Bcl-X(L) protein expression, but appears to be associated with increased expression of cellular Bak and Bax proteins. For more complete data on the mechanisms of action of emodin (9 in total), please visit the HSDB record page. Therapeutic Uses Emodin is a widely used over-the-counter herbal remedy. /EXPL THER:/…Emodin and cassia seed B have been investigated for anticancer applications. Chemoprophylaxis/...These compounds showed significant antitumor promoting effects in a two-stage carcinogenesis study of mouse skin tumors induced by 7,12-dimethylbenzo[a]anthracene as an initiator and 12-O-tetradecanoylphorbol-13-acetate (TPA) as a promoter, whether applied topically or otherwise. Furthermore, emodin showed potent inhibitory activity in a two-stage carcinogenesis study of mouse skin tumors induced by nitric oxide donor (+/-)-(E)-methyl-2-[(E)-hydroxyimino]-5-nitro-6-methoxy-3-hexenamide as an initiator and TPA as a promoter. /EXPL THER:/ ... Emodin improved the adverse effects of high glucose concentrations on human peritoneal mesothelial cells (HPMCs) by inhibiting PKC activation and CREB phosphorylation, suggesting that emodin may have therapeutic potential for preventing or treating glucose-induced peritoneal structural and functional abnormalities. Severe Acute Respiratory Syndrome (SARS) is a newly emerging infectious disease caused by a novel coronavirus (SARS-CoV). The SARS-CoV spike protein (S protein) is a type I membrane-bound protein crucial for the binding of the virus to the host cell receptor angiotensin-converting enzyme 2 (ACE2). We screened 312 Chinese medicinal herbs regulated by the Taiwan Council of Traditional Chinese Medicine and found that three commonly used Polygonaceae herbs could inhibit the interaction between the SARS-CoV S protein and ACE2. The IC50 values of the rhizome of Rheum officinale Baill., the rhizome of Polygonum multiflorum Thunb., and the stem of Polygonum multiflorum ranged from 1 to 10 μg/ml. Emodin, an anthraquinone compound derived from plants in the genera Rheum and Polygonum, significantly blocked the interaction between the S protein and ACE2 in a dose-dependent manner. Furthermore, it inhibited the infection of Vero E6 cells by S protein pseudotyped retroviruses. These findings suggest that emodin may be a potential lead-line drug for the treatment of SARS. [1] Background and Objectives: 11β-hydroxysteroid dehydrogenase type 1 (11β-HSD1) is a highly attractive therapeutic target for type 2 diabetes and metabolic syndrome. Emodin is a natural product and an active ingredient in many traditional Chinese medicines, and has been shown to have various biological activities. This study aimed to investigate the effects of emodin on 11β-HSD1 and its ability to improve metabolic disorders in diet-induced obesity (DIO) mice. Experimental Methods: The inhibitory effect of emodin on recombinant human and mouse 11β-HSD1 was evaluated using scintillation proximity analysis. This study investigated the ability of emodin to inhibit prednisone or dexamethasone-induced insulin resistance in C57BL/6J mice and observed its effects on metabolic abnormalities in diet-induced obesity (DIO) mice. Main Results: Emodin is a potent and selective inhibitor of 11β-HSD1, with IC50 values of 186 nM and 86 nM for human and mouse 11β-HSD1, respectively. A single oral dose of emodin significantly inhibited 11β-HSD1 activity in mouse liver and adipose tissue. Emodin reversed prednisone-induced insulin resistance in mice but had no effect on dexamethasone-induced insulin resistance, confirming its inhibitory effect on 11β-HSD1 in vivo. In DIO mice, oral administration of emodin improved insulin sensitivity and lipid metabolism and reduced mRNA levels of blood glucose, hepatic PEPCK, and glucose-6-phosphatase. Conclusion and significance: This study reveals the novel function of emodin as a potent and selective 11β-HSD1 inhibitor and confirms its benefits for metabolic disorders in DIO mice. This highlights the potential value of emodin analogues as novel compounds for the treatment of metabolic syndrome or type 2 diabetes. [3] Emodin is a natural anthraquinone compound isolated from Polygonaceae plants (e.g., rhubarb, Polygonum cuspidatum) with a variety of pharmacological activities, including antiviral, antitumor, and metabolic regulation effects. [1][2][3] - Antiviral mechanism:Rhein blocks the invasion of SARS-CoV virus by targeting the S protein-ACE2 interaction, which is a key step for the virus to attach to host cells. It does not affect the viral replication mechanism and reduces the risk of drug resistance compared to nucleoside analogs[1] - Antitumor potential: CK2 is overexpressed in a variety of cancers (e.g., breast cancer, colon cancer) and promotes tumor cell proliferation and survival.Rhein selectively inhibits CK2, making it a potential candidate for combination cancer therapy, especially for drugs targeting cell cycle pathways[2] - Metabolic regulation mechanism:Rhein improves insulin resistance and dyslipidemia in DIO mice by inhibiting 11β-HSD1, thereby reducing local cortisol production in the liver and adipose tissue. This avoids systemic glucocorticoid inhibition, which is a limitation of non-selective glucocorticoid inhibitors[3] |
| Molecular Formula |
C15H10O5
|
|
|---|---|---|
| Molecular Weight |
270.24
|
|
| Exact Mass |
270.052
|
|
| Elemental Analysis |
C, 66.67; H, 3.73; O, 29.60
|
|
| CAS # |
518-82-1
|
|
| Related CAS # |
Emodin-d4;132796-52-2
|
|
| PubChem CID |
3220
|
|
| Appearance |
Yellow to orange solid powder
|
|
| Density |
1.6±0.1 g/cm3
|
|
| Boiling Point |
586.9±39.0 °C at 760 mmHg
|
|
| Melting Point |
255 °C (dec.)(lit.)
|
|
| Flash Point |
322.8±23.6 °C
|
|
| Vapour Pressure |
0.0±1.7 mmHg at 25°C
|
|
| Index of Refraction |
1.745
|
|
| LogP |
5.03
|
|
| Hydrogen Bond Donor Count |
3
|
|
| Hydrogen Bond Acceptor Count |
5
|
|
| Rotatable Bond Count |
0
|
|
| Heavy Atom Count |
20
|
|
| Complexity |
434
|
|
| Defined Atom Stereocenter Count |
0
|
|
| SMILES |
CC1=CC2=C(C(=C1)O)C(=O)C3=C(C2=O)C=C(C=C3O)O
|
|
| InChi Key |
RHMXXJGYXNZAPX-UHFFFAOYSA-N
|
|
| InChi Code |
InChI=1S/C15H10O5/c1-6-2-8-12(10(17)3-6)15(20)13-9(14(8)19)4-7(16)5-11(13)18/h2-5,16-18H,1H3
|
|
| Chemical Name |
1,3,8-trihydroxy-6-methylanthracene-9,10-dione
|
|
| Synonyms |
|
|
| HS Tariff Code |
2934.99.9001
|
|
| Storage |
Powder -20°C 3 years 4°C 2 years In solvent -80°C 6 months -20°C 1 month |
|
| Shipping Condition |
Room temperature (This product is stable at ambient temperature for a few days during ordinary shipping and time spent in Customs)
|
| Solubility (In Vitro) |
|
|||
|---|---|---|---|---|
| Solubility (In Vivo) |
Solubility in Formulation 1: 3.33 mg/mL (12.32 mM) in 0.5% CMC-Na/saline water (add these co-solvents sequentially from left to right, and one by one),Suspened solution; with sonication.
Preparation of saline: Dissolve 0.9 g of sodium chloride in 100 mL ddH₂ O to obtain a clear solution. Solubility in Formulation 2: 10 mg/mL (37.00 mM) in 0.5% MC 0.5% Tween-80 (add these co-solvents sequentially from left to right, and one by one), suspension solution; with ultrasonication. (Please use freshly prepared in vivo formulations for optimal results.) |
| Preparing Stock Solutions | 1 mg | 5 mg | 10 mg | |
| 1 mM | 3.7004 mL | 18.5021 mL | 37.0041 mL | |
| 5 mM | 0.7401 mL | 3.7004 mL | 7.4008 mL | |
| 10 mM | 0.3700 mL | 1.8502 mL | 3.7004 mL |
*Note: Please select an appropriate solvent for the preparation of stock solution based on your experiment needs. For most products, DMSO can be used for preparing stock solutions (e.g. 5 mM, 10 mM, or 20 mM concentration); some products with high aqueous solubility may be dissolved in water directly. Solubility information is available at the above Solubility Data section. Once the stock solution is prepared, aliquot it to routine usage volumes and store at -20°C or -80°C. Avoid repeated freeze and thaw cycles.
Calculation results
Working concentration: mg/mL;
Method for preparing DMSO stock solution: mg drug pre-dissolved in μL DMSO (stock solution concentration mg/mL). Please contact us first if the concentration exceeds the DMSO solubility of the batch of drug.
Method for preparing in vivo formulation::Take μL DMSO stock solution, next add μL PEG300, mix and clarify, next addμL Tween 80, mix and clarify, next add μL ddH2O,mix and clarify.
(1) Please be sure that the solution is clear before the addition of next solvent. Dissolution methods like vortex, ultrasound or warming and heat may be used to aid dissolving.
(2) Be sure to add the solvent(s) in order.
| NCT Number | Recruitment | interventions | Conditions | Sponsor/Collaborators | Start Date | Phases |
| NCT00801268 | Terminated | Drug: tripterygium wilfordii Drug: Emodin |
Polycystic Kidney | Zhi-Hong Liu, M.D. | November 2008 | Not Applicable |
 |
|---|